

Volume 27, Issue 2 (Spring 2021)
Intern Med Today 2021, 27(2): 264-287 |
Back to browse issues page
Download citation:
BibTeX | RIS | EndNote | Medlars | ProCite | Reference Manager | RefWorks
Send citation to:



BibTeX | RIS | EndNote | Medlars | ProCite | Reference Manager | RefWorks
Send citation to:
Mardaneh J. Cronobacter Sakazakii: A Foodborne Pathogenic Bacterium in Immunocompromised and Hospitalized Patients. Intern Med Today 2021; 27 (2) :264-287
URL: http://imtj.gmu.ac.ir/article-1-3425-en.html
URL: http://imtj.gmu.ac.ir/article-1-3425-en.html
Department of Microbiology, Infectious Diseases Research Center, School of Medicine, Gonabad University of Medical Sciences, Gonabad, Iran. , jalalmardaneh@yahoo.com
Keywords: Foodborne pathogen, Cronobacter sakazakii, Neonates, Hospitalized patients, Infections, Prevention strategies
Full-Text [PDF 8129 kb]
(1221 Downloads)
| Abstract (HTML) (2634 Views)
Full-Text: (5026 Views)
1. Introduction
History
Cronobacter sakazakii is one of the Enterobacteriaceae family members characterized as a new bacterial species in 1980. Initially, it was identified as an opportunistic pathogen responsible for meningitis and sepsis in neonates [1, 2]. C. sakazakii is of genus Cronobacter and is a genomically heterogeneous, motile (by flagella), and Gram-negative rod bacterium [1]. The bacterium was firstly named as “yellow-pigmented cloacae”, and categorized as “Enterobacter sakazakii” in 1980, considering the diversity in antibiotic susceptibility profiles, pigment production, DNA-DNA hybridization, biochemical reactions, compared with Enterobacter cloacae [3]. By DNA-DNA hybridization method, C. sakazakii has been reported to be 50% associated with Enterobacter cloacae and Citrobacter koseri. Also, C. sakazakii produces brain abscesses similar to those caused by C. koseri [4].
In 1980, Farmer and colleagues determined the species and illustrated 15 biogroups based on the biochemical patterns. A defining characteristic has been the activity of the α-glucosidase enzyme [5]. As a result, selective differential media incorporating chromogenic or fluorogenic α-glucosides such as the indolyl substrate 5-bromo-4-chloro-3-indolyl-α, D-glucopyranoside have been presented [5, 6]. Lately, 16S rDNA sequencing has shown that biochemical diagnostic kits recognized more than one species as C. sakazakii with at least four biochemically and genetically separate subgroups [5, 7, 8].
Nomenclature
Cronobacter sakazakii constitutes a microbiological hazard in the infant food chain, with historically high mortality in neonates. It has been described since early 1929 when it w::as char::acterized by producing a yellow culture and causing septicemia in infants. The name Cronobacter was appropriately derived from Greek mythology. Cronobacter sakazakii comb. nov., was named in honor of the Japanese microbiologist Riichi Sakazaki when the species was first defined in 1980 as Enterobacter sakazakii. Hence, Cronobacter gen. nov. was proposed after the Greek mythological god, Cronus, who swallowed his children at birth [9]. Cronus was the son of Uranus (Heaven) and Gaea (Earth), the youngest of the 12 Titans. He eventually became the king of the Titans and took for his consort his sister Rhea. Rhea bore him several children, including Hestia, Demeter, Hera, Hades, and Poseidon. Cronus, however, had been previously warned by his parents that he would be overthrown by his own child. Then, he swallowed all those children. When Zeus was born, however, Rhea hid him in Crete and tricked Cronus into swallowing a stone instead. Zeus grew up, forced Cronus to disgorge his brothers and sisters, waged war on Cronus, and was victorious [10].
C. sakazakii was previously known as a “yellow-pigmented Enterobacter cloacae” until 1980. It was first classified as a novel species in 1980 when it was introduced as a new species based on differences in biochemical reactions, DNA-DNA hybridization, and antibiotic susceptibility patterns [11-14].
Analysis of both partial 16S rDNA and hsp60 sequences showed that C. sakazakii isolates formed at least four distinct clusters, and it was proposed that clusters 2, 3, and 4 could be unique species. Based on DNA-DNA hybridization and phenotyping, Enterobacter sakazakii was subsequently proposed to be re-classified into a new genus Cronobacter, composed of five distinct species: Cronobacter sakazakii, C. malonaticus, C. turicensis, C. muytjensii, and C. dublinensis. Because of their close relatedness, C. sakazakii and C. malonaticus are difficult to distinguish by 16S rDNA sequence analysis [15, 16].
In 2007, different microorganisms were classified in the Cronobacter genus, including four named species, one unnamed species, and five named subspecies: C. sakazakii subspecies sakazakii, C. sakazakii subspecies malonaticus, C. dublinensis subspecies dublinensis, C. dublinensis subspecies lactaridi, C. dublinensis subspecies lausanensis, C. muytjensii, C. turicensis, C. genomospecies 1 (a distinct species, but unnamed) [17, 18].
Genome and classification
Cronobacter is a Gram-negative member of the Enterobacteriaceae family. It is rod-shaped and motile by peritrichously flagellated. The bacterium was known as Enterobacter sakazakii in 1980, differentiated from Enterobacter cloacae regarding its antibiotic susceptibility profiles, pigment production, biochemical reactions, and DNA-DNA hybridization [19]. Subsequently, Enterobacter sakazakii was identified as a more taxonomically complex and genetically diverse species than earlier thought, and consequently, an extensive poly-phasic survey led to the proposal of the new genus Cronobacter, affording species status to several biogroups of Enterobacter sakazakii [20]. The Cronobacter genus consists of six species: Cronobacter sakazakii, Cronobacter malonaticus, Cronobacter turicensis, Cronobacter muytjensii, Cronobacter dublinensis, and Cronobacter genomospecies 1, to accommodate the biogroups of the hitherto known C. sakazakii [19, 20].
Morphology
Cronobacter sakazakii (ubiquitous and opportunistic pathogen) is a member of the Enterobacteriaceae family. It is a Gram-negative, rod-shaped bacterium measuring approximately between 3 and 1 μm in size and is peritrichously or highly flagellated, thus being motile. It can produce a protective biofilm, too [2, 21].
Metabolism and growth
Strains in the Cronobacter genus are described as catalase positive, oxidase negative, and facultative anaerobic. Substrates used for energy production include amino acids and various lipophilic molecules, sugars, and nitrates in nitrate reduction reactions [22].
Although the optimum temperature for growth of C. sakazakii is 39°C; the temperature range for the growth of Cronobacter is 6°C to 45°C, with an optimal range of 37°C to 43°C. Cronobacter is the most thermo-tolerant member among the Enterobacteriaceae family. Some Cronobacter strains can grow at 47°C and slowly at household refrigeration temperatures. This bacterium can grow below 4°C, indicating that this species would replicate even during refrigeration [23].
Infections caused by Cronobacter sakazakii
Cronobacter sakazakii is a foodborne pathogen associated with the ingestion of IFM that can cause neonatal sepsis, meningitis, and necrotizing enterocolitis. C. sakazakii has been categorized as a severe hazardous bacterium for some individuals, with long duration, substantial chronic sequelae, or life-threatening complications [24].
Cronobacter sakazakii is considered an opportunistic organism and an etiological agent of life-threatening diseases in infant groups. Although the incidence of C. sakazakii infection is low, the prognosis of the disease is poor, and infection is associated with significant morbidity and mortality. Powdered Infant Formula (PIF) milk products contaminated with C. sakazakii have been linked to several clinical cases [3].
The important infections caused by Cronobacter sakazakii in infants are necrotizing enterocolitis, septicemia, and life-threatening meningitis. Low-birth-weight and premature infants and those aged >28 days are more at infection risk than older ones. Clinical manifestations and outcomes include bloodstream infection, necrotizing enterocolitis, meningitis, developmental delay, cyst formation, seizures, brain abscess, hydrocephalus, ventriculitis, and death in 40%–80% of the cases [25-27]. The mortality rate of Necrotizing Enterocolitis (NEC) caused by Cronobacter sakazakii is about 40% to 100% in the most severely affected patients. This infection is the most common gastrointestinal surgical emergency in affected neonates [3].
In neonatal populations, Cronobacter sakazakii infection outbreaks have been related to [ infant formula milk. Most patients who survive C. sakazakii meningitis suffer from severe neurologic sequelae, including development retardation, quadriplegia, and hydrocephalus [28]. Fecal carriage of C. sakazakii has been reported in patients as old as 18 weeks of age, showing the potential for adherence to the mucosal membrane and long-term colonization of the human intestine by this organism and supporting the need for the isolation of infected patients. Neonatal enterocolitis is associated with enteral feeding, intestinal prematurity, and microbial colonization. The infection affects roughly 13% of those with birth weights 1.5 - 2 kg and 5% of all preterm (premature) infants [28, 29].
In children
Cronobacter sakazakii infections continue to be more common in infants and neonates, usually associated with a poor prognosis. Mortality rates of 33% to 80% have been reported [30]. Also, morbidity in C. sakazakii infections is significant. Most children who survive Enterobacter-associated meningitis (94%) develop irreversible neurological sequelae resulting in quadriplegia, developmental delay, and impaired sight and hearing. These complications are frequently attributed to secondary cerebral infarcts. Although premature low-birth-weight infants are at higher risks, C. sakazakii infection has been shown in normal and immunocompromised adults. Some outbreaks of bacterial infection in Neonatal Intensive Care Units (NICUs) have been attributed to powdered formula contaminated with Cronobacter [30, 31].
Infections caused by Cronobacter are mostly associated with sporadic cases of life-threatening diseases, in particular septicemia, NEC, and meningitis in infants. Infants under 28 days of age and neonates with low birth weight (i.e., < 2.5 kg) are at heightened risk compared to more mature counterparts [19]. Various infections include bloodstream infections, NEC characterized by intestinal necrosis and pneumatosis intestinalis, meningitis leading to ventriculitis, hydrocephalus, brain abscess, cyst formation, pulmonary and urinary tract infections. In infants, meningitis caused by Cronobacter is established 4 or 5 days after birth and can be fatal within hours to several days following the onset of first clinical signs. The mortality rate for neonatal infections has been reported to be as high as 80%, and survivors often suffer from severe irreversible neurological disorders [19, 32].
In adults
There have been few reports of C. sakazakii infection in adults, and it is not usually life-threatening.
Mortality and morbidity
The implications for infected neonates are severe. Once neonates are infected, they have a mortality rate of 40% to 80%, and a 20% chance of survival is accompanied by serious neurological complications [33, 34].
Epidemiology and reservoirs of Cronobacter sakazakii
Given the ubiquity of Cronobacter sakazakii animate environment (animals, man) and inanimate environment (plants, soil, water), it is not surprising that C. sakazakii is detected in many foods and food products of animal and vegetable origin [35].
Cronobacter sakazakii has been isolated from the products, such as chocolate, cereal, pasta, potato flour, milk powder, and spices. Also, this bacterium has been recovered from the guts of the stable fly (Stomoxys calcitrans), the Mexican fruit fly (Anastrpha ludens), and household vacuum cleaner bags [26, 36]. Oral and intestinal colonization with C. sakazakii may be associated with the ingestion of contaminated foods. As C. sakazakii is an opportunistic pathogen, the patient’s colonization flora is the most probable source of infection under circumstances of immunosuppression and severe underlying diseases in patients after the neonatal period [35, 37]. C. sakazakii has been isolated from plant food and food ingredients like cereal, fruit and vegetables, legume products, herbs and spices as well as animal food sources like milk, meat, and fish and products made from these foods. The spectrum of C. sakazakii-contaminated food covers both raw and processed foods [35]. Food and food ingredients may be contaminated with C. sakazakii under hygiene mismanagement by contaminated insects and rats. C. sakazakii has been detected in food production as well as in domestic environments. The ubiquitous microorganism C. sakazakii has been isolated from a wide spectrum of environmental sources, including water, waste and thermal spring water, soil, dust from households, and food production-lines [3, 35, 38, 39].
C. sakazakii-contaminated vegetarian food and drinks comprise vegetables, fruit, legumes, cereal, herbs, spices, and other related products. C. sakazakii has been isolated from the xylem fluid of lemon rootstocks, the rhizosphere of wheat, and as endophytic bacteria from the leaves of rice plants [35]. It has also been detected in the bacterial colonization flora of disinfected sugar beet seeds. As C. sakazakii belongs to the cultivable endophytic and epiphytic flora of rice and soybean plants, it could be isolated from the related food products [35, 40, 41]. Some traditional cereal-, herb- and legume-based food and beverages were contaminated with C. sakazakii. The bacterium may be part of starter cultures for the fermentation of traditional vegetarian food products. C. sakazakii has been detected in mixed salad vegetables and imported, fresh, and deep-frozen vegetables at the retail level [35, 42].
C. sakazakii-contaminated food of animal origin comprise a variety of meat and meat products from camel, pig, beef, and poultry, besides eggs, raw milk and different dairy products and, less frequently, fish [35]. C. sakazakii has been isolated from animals, especially from birds, lizards, rats, and piglets. In vertebrates, C. sakazakii is a member of the normal (animal and human) oral and intestinal flora. It was found among the isolates from secretions of an infected mammary gland of dairy heifers. Liu et al. detected C. sakazakii in feed for pets. Also, C. sakazakii was isolated from various raw and ready-to-eat meat (products) [35, 43]. Isolated C. sakazakii has been found during a complicated curing process of meat products. It is a histamine-forming microorganism in the ripening process of cheese and has been isolated from a cheese whey substrate. Researchers demonstrated the lipolytic activity of a C. sakazakii-strain. C. sakazakii has been detected in fresh and prepared fish, too. A tetracycline-resistant C. sakazakii-strain from a Chilean freshwater salmon farm with no history of recent antibiotic use has been isolated. It has also been isolated from smoked sardines after 12 weeks of storage after irradiation [35, 44].
C. sakazakii with the total frequencies of 1.8% (10/564 strains) and 0.4% (1/256 strains) was detected when investigating the central and local drinking water supplies, respectively. During their investigation for biofilm formation, the bacteria indigenous to the water distribution system was found. Even bottled beverages should not be considered free of microorganisms [35, 45].
Transmission
Cronobacter sakazakii is thermo-tolerant and can contaminate PIF both intrinsically and extrinsically. Intrinsic contamination results from introducing the organism into the PIF at some stages during the manufacturing process. In contrast, extrinsic contamination may result from the use of contaminated utensils, such as blenders and spoons, to prepare PIF [46].
C. sakazakii are known to contaminate infant formula through the raw materials used for producing the formula or other dry ingredients after pasteurization; the caregiver may reconstitute the contamination of the formula just before feeding [47]. In most cases, the transmission route of Cronobacter-associated infant infections has not been confirmed. However, epidemiological studies in several incidents have implicated rehydrated PIF products as the vehicle of transmission and utensils used for formulae preparation as the likely source of contamination [19, 48].
Infection incidence
Although the number of well-documented cases of Cronobacter sakazakii infections in infants worldwide has increased in recent years, it has remained very low compared to many other infectious diseases. Also, there are reports in the literature about C. sakazakii (Cronobacter spp.) infections in older age groups, including adults, but the rate of infection among infants is lower. There has been no reported systematic review of cases in children and adults [49].
Pathogenicity and virulent factors
Virulence determinants of Cronobacter sakazakii are still unknown, and pathogenicity mechanisms have only begun to be researched. C. sakazakii is a foodborne pathogen that can cause severe illnesses and even death in persons with immunological deficiencies such as neonates, hospitalized patients in NICU, persons with severe underlying diseases, and the elderly. In these populations, this bacterium can successfully establish, colonize, and ultimately produce severe diseases [50, 51]. In humans, C. sakazakii has been found to affect specifically the nervous, gastrointestinal, and vascular systems. Establishment in the human vascular system causes bacteremia and or sepsis. It is often preceded by colonization beyond the blood-brain barrier leading to cerebrospinal fluid infection and meningitis, which may progress into intracerebral infarctions, brain abscess, and or cyst formation resulting in central nervous system deterioration. Necrotizing Enterocolitis (NEC) caused by C. sakazakii is currently the most common gastrointestinal emergency in neonates, characterized by necrosis of the gastrointestinal lumen [50-52].
Adhesions and attachment
Cronobacter sakazakii has adhesive capacities to several in vitro cell lines, including endothelial and transformed epithelial lines. This bacterium can attach to intestinal cells and survive in macrophages, but the specific involved receptors have remained to be determined. Recently, it was shown that the disruption of tight junctions significantly enhances the association of C. sakazakii with Caco-2 cells [53]. Some reports suggest a similarity between the tropism of Cronobacter sakazakii and Citrobacter koseri for invasion and infection of the central nervous system. It was noted that brain abscesses due to Cronobacter and Citrobacter koseri were morphologically similar and may be due to similar virulence mechanisms [53, 54].
Cronobacter sakazakii carries endotoxin on its surface; however, other virulence factors may also be crucial to pathogenicity. For a pathogen to cause an infection in a host, it should adhere to and colonize host surfaces. Specific adhesion to host cells is considered to be an essential virulence factor for most bacterial pathogens. This foodborne pathogen appears to adhere instantaneously to host surfaces and then proliferates in a logarithmic manner until an optimal concentration is attained. The adhesion of C.sakazakii to epithelial cells is mainly non-fimbriae-based, suggesting the role of other virulence factors in binding. The bacterium exhibits clustered adhesion, a pattern that also has been associated with neonatal colitis-causing strains of Klebsiella pneumoniae [28, 32]. C. sakazakii has an unusual surviving ability under dry conditions, but the thermal tolerance of C. sakazakii-strains may be different [32, 35].
Exposure to Cronobacter induced apoptosis and increased expression of interleukin-6 in infant rats could effectively explain the pathogenicity of Cronobacter-associated NEC in neonates [19].
The adhesive behavior of C. sakazakii strains is impressive, but until only recently, it has been thoroughly investigated. Adherence has been reported on inorganic and synthetic surfaces, such as plastics, silicon, PVC, polycarbonate, glass, stainless steel, and public water systems. Adhesion to living surfaces such as in a pathogenic capacity has been shown to differ between strains, but all share the trait of being independent of fimbriae structures and require a metabolically active host cell for adhesion [55].
Capsule
Some strains of Cronobacter sakazakii produce a viscous capsular material, potentially allowing the organism to form a biofilm on feeding equipment and contact surfaces, and how this material contributes to macrophage evasion remains to be determined [50, 56]. C. sakazakii capsule may also protect the organism, facilitating its survival in desiccated environments. The biofilm and capsule have been shown to reduce the efficacy of various common sanitizing methods, such as UV-light radiation, high osmotic pressures, heat, dry conditions, starvation, low pH, detergents, antibiotics, phagocytes, antibodies, and some bacteriophages. This biofilm may protect C. sakazakii, allowing it to survive osmotic, thermal, and tensile stressors [56]. Enterobacter sakazakii also produces a viscous capsular material, potentially allowing the organism to form a biofilm on feeding equipment and contact surfaces [38]. This biofilm may protect ES, allowing it to survive osmotic, thermal, and tensile stressors. Mange et al. demonstrated that ES has adhesive capacities to several in vitro cell lines, including endothelial and transformed epithelial lines [19].
Furthermore, entry is enhanced in the presence of the host cell cytoskeleton (actin filaments and microtubule structures) and disruption of the tight junction. This event may account for the opportunistic nature of this infection, particularly in infants and neonates [19].
The abilities of C. sakazakii to produce capsule and biofilm reduce the efficacy of UV-light, high osmotic pressure, heat, dry conditions, starvation, acids, inhibitory detergents like sanitizers and antibiotics, phagocytes, antibodies, and bacteriophages. C. sakazakii-strains can adhere to surfaces of materials and are used in the production, preparation, and administration of food like plastics, silicon, latex, polyvinyl chloride, less frequently on glass and stainless steel. C. sakazakii has been cultivated from biofilms of drinking water systems [35].
Outer membrane protein A (ompA)
Cronobacter sakazakii expresses the outer membrane protein A (ompA) that shows a high degree of homology with ompA genes of other Gram-negative bacteria [28]. The ompA protein is crucial in the invasion of brain endothelial cells by C. sakazakii. The organism also induces microtubule condensation at the sites of entry in endothelial cells. OmpA expression is required for moderate invasion of human intestinal epithelial cells. The binding of this bacterium to enterocytes, both in vitro and in the animal model, induces enterocyte apoptosis/necrosis in a dose-dependent manner [57].
Also, Cronobacter invades human brain microvascular endothelial cells more often than epithelial cells, but entry requires the expression of outer membrane protein A (ompA) to induce microtubule condensation [19].
Biofilm formation
Cronobacter sakazakii can attach to plastics and silicon rubber surfaces and grow in a biofilm. Enteral feeding tubes and feeding-bottle teats can harbor the bacterium in large numbers. Biofilm formation may also be a factor associated with altered susceptibility to antimicrobials [3].
Lipopolysaccharide
Large amounts of Lipopolysaccharide (LPS) are released by the bacteria killed by antibiotics, phagocytosis, the complement complex, or treatment with divalent cation chelators. In an infected host, small amounts of LPS can be protective by stimulating the immune system, while large amounts induce high fever and lead to septic shock and death by multiorgan failure and systemic inflammatory response. LPS liberated from bacteria is associated with LPS Binding Protein (LBP), an acute-phase protein present in the bloodstream, and forms complexes consisting of LPS, LBP, and soluble CD14 (sCD14) [58, 59]. The O-antigen region is one of the most variable regions in the membranes of Gram-negative bacteria, and this variability is routinely used to discriminate serotypes within the species of bacteria. Within O-antigen gene clusters, the wzx and wzy genes have the most diverse nucleotide sequences, which has led to their wide use in serotype-specific PCR assays [59, 60]. Finally, high levels of heat-stable lipopolysaccharide (endotoxin) in infant formula may enhance the translocation of Cronobacter across the gut and blood-brain barrier, increasing the risk of bacteremia in neonates [19, 58].
Viable but Nonculturable state (the VNC state)
Cronobacter sakazakii can become injured when they are subjected to stress such as heat and drying. This bacterium can respond to stress by entering a unique physiological state known as the “viable but nonculturable” (often abbreviated as the VNC state). Bacterial pathogens in the Enterobacteriaceae family (for example, Escherichia coli O157:H7, Salmonella, Shigella, Enterobacter cloacae, Klebsiella) can enter this state [17].
Minimum lethal dose
The actual amount of ES contamination usually is low, ranging from 0.36 to 66 Colony-Forming Units (CFU)/100 g [28].
Immune system state
In industrial countries, persons may have a predisposing condition (e.g. those receiving chemotherapy, transplant persons, people with chronic diseases, HIV positive patients, those receiving immunosuppressive treatments) that suppresses their T-cell mediated immunity, thereby potentially raising their susceptibility to a broad range of bacterial infections, including Cronobacter sakazakii infections. Also, an unknown proportion of neonates, infants, and toddlers may be temporarily immunocompromised at any time because of the pharmaceutical treatments, such as steroids for asthma exacerbations, injury, effects of acute disease, or stress. In addition, there has been a significant increase in the prescription of gastric acid-suppressing medications for gastroesophageal reflux in infants and toddlers in industrialized settings [61, 62]. While not traditionally considered immuno-suppressing, such medications impair one of the first lines of defense humans have against ingested pathogens. Infants, in addition, have lower levels of gastric acid, making them more vulnerable to infections. The prevalence of immunocompromising conditions may be relatively low in the developed countries, but the picture seems very different in developing countries, where the prevalence of such factors can rise to 40% [62].
Cronobacter sakazakii in powdered infant formula milk
Some of the vehicles of transmission and the infectious dose of Cronobacter sakazakii are unknown. The number of reported cases of Cronobacter infections is very low; nevertheless, it has slightly increased recently. Cronobacter sakazakii does not withstand the temperature at which milk is pasteurized, but it is easily found post-pasteurization, manufacturer environment, and preparation and handling before consumption. The thermal tolerance of Cronobacter in dried PIF has been well documented. This is why important steps towards preventive measures are needed to eliminate or avoid risks of PIF contamination and proliferation of the C. sakazakii. It is worth pointing out that breastfeeding should always be supported and encouraged since breast milk constitutes the preferred food for newborn infants, especially in their early months. When this is not possible, a mother should be well informed and educated on the importance of hygiene while handling, preparing, and storing PIF. Recommendations also underline the importance of using ready-to-feed PIF, complying with the rules for aseptic preparation and refrigeration at 2°C to 3°C of reconstituted PIF for a span of time shorter than 4 hours [2].
Treatment of infection caused by Cronobacter sakazakii
Traditionally, the treatment of Cronobacter infections with a combination of β-lactam (ampicillin) and aminoglycoside (gentamycin) or ampicillin-chloramphenicol has been successful [19]. It is sensitive to some antibiotics such as numerous β-lactams, antifolates, tetracycline, aminoglycosides, chloramphenicol, and quinolones. Also, trimethoprim-sulfamethoxazole may be useful. Nevertheless, many Cronobacter species are resistant to narrow-spectrum penicillins (i.e., ampicillin and amoxicillin) that traditionally have had good activity against members of the Enterobacteriaceae family. Emerging resistance of this bacterium to some antibiotics of choice should prompt physicians to consider carbapenems (ertapenem, meropenem, imipenem) or the newer broad-spectrum cephalosporins (e.g. cefepime) in combination with a second agent such as an aminoglycoside (e.g. gentamycin). Consequently, selecting antibiotics based on bacterial pure culture and susceptibility results and minimizing the use of broad-spectrum drugs are of paramount importance. C. sakazakii has intrinsic resistance to fusidic acids, streptogramins, glycopeptides, and lincosamides [28, 63].
Antimicrobial Resistance of Cronobacter sakazakii
C. sakazakii is intrinsically resistant to fosfomycin, rifampicin (rifampin), lincomycin, clindamycin, macrolides (clarithromycin, erythromycin, azithromycin), fusidic acid, and streptogramins. Nevertheless, resistance to ampicillin has increased owing to the production of β-lactamases and the acquisition of Transposable Elements (TEs) or jumping genes [2, 3, 64, 65]. Cronobacter sakazakii can inactivate cephalosporins and broad-spectrum antibiotics by secretion of β-lactamase enzymes [3, 64, 65].
Cronobacter sakazakii Resistance to routine sterilization methods
Cronobacter sakazakii is relatively resistant to dryness, heat, and osmotic stresses, which may explain, in part, its presence and survival in desiccated infant powder and similarly prepared products. Unsuitable storage and temperature regulation may raise the bacterial load, consequently facilitating infection outbreaks. Although C. sakazakii infection may increase secondary to reheating of formula and poor storage, it has not been documented definitely that staff of hospital themselves are not a vector for this foodborne pathogen. Unfortunately, hospital personnel may contribute to the spread of infection by ignoring suitable hygiene guidelines [66]. So, great care should be taken in contact with the isolation of infected and susceptible patients (infants) and adhering to strict hand washing. Also, C. sakazakii may form biofilms on different surfaces and, as a result of that acquiring resistance to disinfectants. It can survive in food powders for at least 12 months [28, 66, 67].
Tolerance or resistance of desiccated conditions for such a long time can be attributed to certain aspects of the Cronobacter sakazakii physiology, perhaps most notably because some isolates can produce a polysaccharide capsule. Over a 12-month storage period, the organism has been shown to survive better in dried formula products with an aw between 0.25 and 0.30 than at 0.69 and 0.82. C. sakazakii is resistant to desiccation over a wide range of aw (0.25-0.86) [68]. Sanitizer, disinfectants, and detergents reduce the C. sakazakii thermal resistance in powdered infant formula milk and other nutrient products.
Laboratory detection of Cronobacter sakazakii
An obstacle in diagnosing C. sakazakii infections and their sources is the ability or capacity of clinical, food, and environmental laboratories to identify the organism. The isolation of bacteria, such as C. sakazakii, from dried foods requires a series of steps to revitalize stressed cells that would otherwise not be cultured.
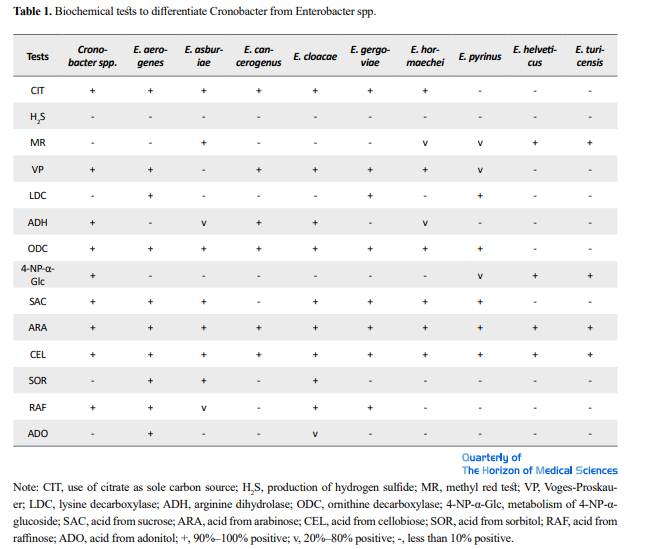
Cronobacter is a genus of the Enterobacteriaceae family. It is a Gram-negative, facultatively anaerobic, oxidase-negative, catalase-positive, and rod-shaped bacterium. They are generally motile, reduce nitrate, use citrate, hydrolyze esculin and arginine, and are positive for L-ornithine decarboxylation. Acid is produced from D-glucose, D-sucrose, D-raffinose, D-melibiose, D-cellobiose, D-mannitol, D-mannose, L-rhamnose, L-arabinose, D-trehalose, galacturonate and D-maltose. Cronobacter spp. are also generally positive for acetoin production (Voges–Proskauer test) and negative for the methyl red test, indicating 2,3-butanediol rather than mixed acid fermentation [68-71]. Some important biochemical tests to differentiate Cronobacter from Enterobacter spp. are presented in Table 1.
Isolation and identification from clinical samples
The detection of bacteria from typically sterile sites (e.g. blood, CSF) is less complex than their detection and isolation from a powdered formula, as the organisms would not be stressed and are unlikely to be in a mixed population. Nevertheless, accurate clinical identification of isolates such as C. sakazakii is restricted by using methods that have not been specifically validated for the organism [70].
Isolation and identification from food
Isolation of Cronobacter sakazakii from food samples is performed according to FDA protocol. Three Erlenmeyer flasks of sterile distilled water (pre-warmed to 45°C) of 9, 90, and 900 mL containing 1, 10, and 100 g of PIF, respectively, are prepared. After the weighed samples are thoroughly mixed and dissolved in distilled water, it was incubated at 35±2°C for 18 to 24 hours. After incubation, 10 mL of each sample was added to 90 mL of Enterobacteriaceae Enrichment (EE) broth medium and placed at 35±2°C for 18 to 24 hours. Following the incubation, a loopful of the enrichment culture is streaked onto duplicate Violet Red Bile Glucose Agar (VRBGA) medium plates, and incubation was performed for 18 to 24 hours at 35°C±2°C. Four presumptive colonies are picked from each VRBGA plate, and pure culture was performed on MacConkey agar and Tryptic Soy Agar (TSA). The isolated colonies that produce yellow pigment on TSA medium at room temperature (25°C) are identified. For the final confirmation of isolates, manual biochemical tests and biochemical tests embedded in the API-20E biochemical kit system are used [2].
Different media, including E. sakazakii Selective Broth (ESSB), Druggan-Forsythe-Iversen agar, Cronobacter Screening Broth (CSB), E. sakazakii Enrichment Broth (ESE), EE broth, and modified Lauryl Sulfate Broth (mLST), are used for the isolation of C. sakazakii from food samples. Compared with conventional, labor-intensive methods, the recently developed fluoro- and chromo-genic differential selective media decrease the time for the isolation of C. sakazakii [72]. The phenotypic identification of C. sakazakii with commercial systems may be inadequate for the strains isolated from food specimens. Molecular genetic methods revealed that several strains identified as C. sakazakii by commercial biochemical kits belonged to distinct species. To confirm C. sakazakii, more than one differentiation system is recommended. Computationally-based methods, including biochemical and 16S rDNA-data, improve the reliability and the detection time of identification [35, 73].
Molecular detection of Cronobacter sakazakii
Different techniques have been designed to identify Cronobacter sakazakii. Key molecular tests and particular areas of the DNA sequence can be applied for the accurate discrimination of C. sakazakii from other closely related species. To date, assays for the isolation and identification of C. sakazakii have applied production of yellow pigment and the α-glucosidase test as presumptive differentiating characteristics. However, these assays can result in presumptive false positives because of groups of as yet unidentified non-C. sakazakii Enterobacteriaceae, which are also positive for both of these characteristics. The use of yellow pigment as a defining characteristic can also result in false-negative because of the occurrence of non-pigmented C. sakazakii isolates and the occasional transient nature of this trait [5].
Prevention strategies
There are no active surveillance systems and guidelines for Cronobacter sakazakii disease. The World Health Organization and Food and Agriculture Organization of the United Nations made the following recommendations: (1) notify infant caregivers of the risks related to non-sterile powdered formula products, (2) encourage industry partners to develop a range of affordable sterile formula options, (3) if infants cannot breastfeed, consider feeding high-risk infants with sterile formula, and (4) consider setting an industry guideline for C. sakazakii and other Enterobacteriaceae members in infant formula milk. Factories warning labels on packages of powdered infant formula should stress that their products are non-sterile and need suitable handling, preparation, and storage and that sterile, liquid formula alternatives are available [26].
Pasteurization is effective in destroying Cronobacter sakazakii. Acidification reduced the concentration of C. sakazakii in vegetable-based foods and different types of infant formula products [35, 42]. Hygiene mismanagement due to incorrect time and temperature factors due to the transmission of bacteria via small vertebrates, insects, hands, and equipment should be avoided during the preparation process, production, and storage of food and drink. C. sakazakii may be related to food spoilage. It should be noted that the detection of the ubiquitous C. sakazakii in food is not always an indicator of hygiene mismanagement. For persons with acquired or congenital immunological deficiency, especially neonates, infants, elderly individuals, and persons with severe underlying diseases, the widespread occurrence of C. sakazakii and other Enterobacteriaceae members in the food and environment may indicate a health hazard [35].
2. Conclusion
Because of the ubiquitous nature of Cronobacter sakazakii and its unclear pathogenesis, preventive measures by parents, infant formula manufacturers, and health care providers will be necessary for the prevention of C. sakazakii-related infections. We recommend focusing on simple preventative strategies such as the promotion of breast milk feeding, the inclusion of warnings on powdered infant formula packages that they may be contaminated with C. sakazakii, and abstinence from the practice of re-warming of reconstituted formula. In adults, we believe that reconstituted dairy products should be avoided in immunosuppressed populations. Appropriate barrier precautions should be observed in ICU settings (both adult and neonatal), where the spread of infection may be more prevalent.
Ethical Considerations
Compliance with ethical guidelines
This article is a meta-analysis with no human or animal sample.
Funding
This study was supported financially by Gonabad University of Medical Sciences, Gonabad City, Iran.
Authors' contributions
All authors listed have contributed sufficiently to the project to be included as authors, and all those who are qualified to be authors are listed in the author byline.
Conflicts of interest
All authors declared no conflict of interest.
Acknowledgements
Our thanks go to Hassan Khajehei, PhD, for copy editing of the manuscript.
Refrences:
History
Cronobacter sakazakii is one of the Enterobacteriaceae family members characterized as a new bacterial species in 1980. Initially, it was identified as an opportunistic pathogen responsible for meningitis and sepsis in neonates [1, 2]. C. sakazakii is of genus Cronobacter and is a genomically heterogeneous, motile (by flagella), and Gram-negative rod bacterium [1]. The bacterium was firstly named as “yellow-pigmented cloacae”, and categorized as “Enterobacter sakazakii” in 1980, considering the diversity in antibiotic susceptibility profiles, pigment production, DNA-DNA hybridization, biochemical reactions, compared with Enterobacter cloacae [3]. By DNA-DNA hybridization method, C. sakazakii has been reported to be 50% associated with Enterobacter cloacae and Citrobacter koseri. Also, C. sakazakii produces brain abscesses similar to those caused by C. koseri [4].
In 1980, Farmer and colleagues determined the species and illustrated 15 biogroups based on the biochemical patterns. A defining characteristic has been the activity of the α-glucosidase enzyme [5]. As a result, selective differential media incorporating chromogenic or fluorogenic α-glucosides such as the indolyl substrate 5-bromo-4-chloro-3-indolyl-α, D-glucopyranoside have been presented [5, 6]. Lately, 16S rDNA sequencing has shown that biochemical diagnostic kits recognized more than one species as C. sakazakii with at least four biochemically and genetically separate subgroups [5, 7, 8].
Nomenclature
Cronobacter sakazakii constitutes a microbiological hazard in the infant food chain, with historically high mortality in neonates. It has been described since early 1929 when it w::as char::acterized by producing a yellow culture and causing septicemia in infants. The name Cronobacter was appropriately derived from Greek mythology. Cronobacter sakazakii comb. nov., was named in honor of the Japanese microbiologist Riichi Sakazaki when the species was first defined in 1980 as Enterobacter sakazakii. Hence, Cronobacter gen. nov. was proposed after the Greek mythological god, Cronus, who swallowed his children at birth [9]. Cronus was the son of Uranus (Heaven) and Gaea (Earth), the youngest of the 12 Titans. He eventually became the king of the Titans and took for his consort his sister Rhea. Rhea bore him several children, including Hestia, Demeter, Hera, Hades, and Poseidon. Cronus, however, had been previously warned by his parents that he would be overthrown by his own child. Then, he swallowed all those children. When Zeus was born, however, Rhea hid him in Crete and tricked Cronus into swallowing a stone instead. Zeus grew up, forced Cronus to disgorge his brothers and sisters, waged war on Cronus, and was victorious [10].
C. sakazakii was previously known as a “yellow-pigmented Enterobacter cloacae” until 1980. It was first classified as a novel species in 1980 when it was introduced as a new species based on differences in biochemical reactions, DNA-DNA hybridization, and antibiotic susceptibility patterns [11-14].
Analysis of both partial 16S rDNA and hsp60 sequences showed that C. sakazakii isolates formed at least four distinct clusters, and it was proposed that clusters 2, 3, and 4 could be unique species. Based on DNA-DNA hybridization and phenotyping, Enterobacter sakazakii was subsequently proposed to be re-classified into a new genus Cronobacter, composed of five distinct species: Cronobacter sakazakii, C. malonaticus, C. turicensis, C. muytjensii, and C. dublinensis. Because of their close relatedness, C. sakazakii and C. malonaticus are difficult to distinguish by 16S rDNA sequence analysis [15, 16].
In 2007, different microorganisms were classified in the Cronobacter genus, including four named species, one unnamed species, and five named subspecies: C. sakazakii subspecies sakazakii, C. sakazakii subspecies malonaticus, C. dublinensis subspecies dublinensis, C. dublinensis subspecies lactaridi, C. dublinensis subspecies lausanensis, C. muytjensii, C. turicensis, C. genomospecies 1 (a distinct species, but unnamed) [17, 18].
Genome and classification
Cronobacter is a Gram-negative member of the Enterobacteriaceae family. It is rod-shaped and motile by peritrichously flagellated. The bacterium was known as Enterobacter sakazakii in 1980, differentiated from Enterobacter cloacae regarding its antibiotic susceptibility profiles, pigment production, biochemical reactions, and DNA-DNA hybridization [19]. Subsequently, Enterobacter sakazakii was identified as a more taxonomically complex and genetically diverse species than earlier thought, and consequently, an extensive poly-phasic survey led to the proposal of the new genus Cronobacter, affording species status to several biogroups of Enterobacter sakazakii [20]. The Cronobacter genus consists of six species: Cronobacter sakazakii, Cronobacter malonaticus, Cronobacter turicensis, Cronobacter muytjensii, Cronobacter dublinensis, and Cronobacter genomospecies 1, to accommodate the biogroups of the hitherto known C. sakazakii [19, 20].
Morphology
Cronobacter sakazakii (ubiquitous and opportunistic pathogen) is a member of the Enterobacteriaceae family. It is a Gram-negative, rod-shaped bacterium measuring approximately between 3 and 1 μm in size and is peritrichously or highly flagellated, thus being motile. It can produce a protective biofilm, too [2, 21].
Metabolism and growth
Strains in the Cronobacter genus are described as catalase positive, oxidase negative, and facultative anaerobic. Substrates used for energy production include amino acids and various lipophilic molecules, sugars, and nitrates in nitrate reduction reactions [22].
Although the optimum temperature for growth of C. sakazakii is 39°C; the temperature range for the growth of Cronobacter is 6°C to 45°C, with an optimal range of 37°C to 43°C. Cronobacter is the most thermo-tolerant member among the Enterobacteriaceae family. Some Cronobacter strains can grow at 47°C and slowly at household refrigeration temperatures. This bacterium can grow below 4°C, indicating that this species would replicate even during refrigeration [23].
Infections caused by Cronobacter sakazakii
Cronobacter sakazakii is a foodborne pathogen associated with the ingestion of IFM that can cause neonatal sepsis, meningitis, and necrotizing enterocolitis. C. sakazakii has been categorized as a severe hazardous bacterium for some individuals, with long duration, substantial chronic sequelae, or life-threatening complications [24].
Cronobacter sakazakii is considered an opportunistic organism and an etiological agent of life-threatening diseases in infant groups. Although the incidence of C. sakazakii infection is low, the prognosis of the disease is poor, and infection is associated with significant morbidity and mortality. Powdered Infant Formula (PIF) milk products contaminated with C. sakazakii have been linked to several clinical cases [3].
The important infections caused by Cronobacter sakazakii in infants are necrotizing enterocolitis, septicemia, and life-threatening meningitis. Low-birth-weight and premature infants and those aged >28 days are more at infection risk than older ones. Clinical manifestations and outcomes include bloodstream infection, necrotizing enterocolitis, meningitis, developmental delay, cyst formation, seizures, brain abscess, hydrocephalus, ventriculitis, and death in 40%–80% of the cases [25-27]. The mortality rate of Necrotizing Enterocolitis (NEC) caused by Cronobacter sakazakii is about 40% to 100% in the most severely affected patients. This infection is the most common gastrointestinal surgical emergency in affected neonates [3].
In neonatal populations, Cronobacter sakazakii infection outbreaks have been related to [ infant formula milk. Most patients who survive C. sakazakii meningitis suffer from severe neurologic sequelae, including development retardation, quadriplegia, and hydrocephalus [28]. Fecal carriage of C. sakazakii has been reported in patients as old as 18 weeks of age, showing the potential for adherence to the mucosal membrane and long-term colonization of the human intestine by this organism and supporting the need for the isolation of infected patients. Neonatal enterocolitis is associated with enteral feeding, intestinal prematurity, and microbial colonization. The infection affects roughly 13% of those with birth weights 1.5 - 2 kg and 5% of all preterm (premature) infants [28, 29].
In children
Cronobacter sakazakii infections continue to be more common in infants and neonates, usually associated with a poor prognosis. Mortality rates of 33% to 80% have been reported [30]. Also, morbidity in C. sakazakii infections is significant. Most children who survive Enterobacter-associated meningitis (94%) develop irreversible neurological sequelae resulting in quadriplegia, developmental delay, and impaired sight and hearing. These complications are frequently attributed to secondary cerebral infarcts. Although premature low-birth-weight infants are at higher risks, C. sakazakii infection has been shown in normal and immunocompromised adults. Some outbreaks of bacterial infection in Neonatal Intensive Care Units (NICUs) have been attributed to powdered formula contaminated with Cronobacter [30, 31].
Infections caused by Cronobacter are mostly associated with sporadic cases of life-threatening diseases, in particular septicemia, NEC, and meningitis in infants. Infants under 28 days of age and neonates with low birth weight (i.e., < 2.5 kg) are at heightened risk compared to more mature counterparts [19]. Various infections include bloodstream infections, NEC characterized by intestinal necrosis and pneumatosis intestinalis, meningitis leading to ventriculitis, hydrocephalus, brain abscess, cyst formation, pulmonary and urinary tract infections. In infants, meningitis caused by Cronobacter is established 4 or 5 days after birth and can be fatal within hours to several days following the onset of first clinical signs. The mortality rate for neonatal infections has been reported to be as high as 80%, and survivors often suffer from severe irreversible neurological disorders [19, 32].
In adults
There have been few reports of C. sakazakii infection in adults, and it is not usually life-threatening.
Mortality and morbidity
The implications for infected neonates are severe. Once neonates are infected, they have a mortality rate of 40% to 80%, and a 20% chance of survival is accompanied by serious neurological complications [33, 34].
Epidemiology and reservoirs of Cronobacter sakazakii
Given the ubiquity of Cronobacter sakazakii animate environment (animals, man) and inanimate environment (plants, soil, water), it is not surprising that C. sakazakii is detected in many foods and food products of animal and vegetable origin [35].
Cronobacter sakazakii has been isolated from the products, such as chocolate, cereal, pasta, potato flour, milk powder, and spices. Also, this bacterium has been recovered from the guts of the stable fly (Stomoxys calcitrans), the Mexican fruit fly (Anastrpha ludens), and household vacuum cleaner bags [26, 36]. Oral and intestinal colonization with C. sakazakii may be associated with the ingestion of contaminated foods. As C. sakazakii is an opportunistic pathogen, the patient’s colonization flora is the most probable source of infection under circumstances of immunosuppression and severe underlying diseases in patients after the neonatal period [35, 37]. C. sakazakii has been isolated from plant food and food ingredients like cereal, fruit and vegetables, legume products, herbs and spices as well as animal food sources like milk, meat, and fish and products made from these foods. The spectrum of C. sakazakii-contaminated food covers both raw and processed foods [35]. Food and food ingredients may be contaminated with C. sakazakii under hygiene mismanagement by contaminated insects and rats. C. sakazakii has been detected in food production as well as in domestic environments. The ubiquitous microorganism C. sakazakii has been isolated from a wide spectrum of environmental sources, including water, waste and thermal spring water, soil, dust from households, and food production-lines [3, 35, 38, 39].
C. sakazakii-contaminated vegetarian food and drinks comprise vegetables, fruit, legumes, cereal, herbs, spices, and other related products. C. sakazakii has been isolated from the xylem fluid of lemon rootstocks, the rhizosphere of wheat, and as endophytic bacteria from the leaves of rice plants [35]. It has also been detected in the bacterial colonization flora of disinfected sugar beet seeds. As C. sakazakii belongs to the cultivable endophytic and epiphytic flora of rice and soybean plants, it could be isolated from the related food products [35, 40, 41]. Some traditional cereal-, herb- and legume-based food and beverages were contaminated with C. sakazakii. The bacterium may be part of starter cultures for the fermentation of traditional vegetarian food products. C. sakazakii has been detected in mixed salad vegetables and imported, fresh, and deep-frozen vegetables at the retail level [35, 42].
C. sakazakii-contaminated food of animal origin comprise a variety of meat and meat products from camel, pig, beef, and poultry, besides eggs, raw milk and different dairy products and, less frequently, fish [35]. C. sakazakii has been isolated from animals, especially from birds, lizards, rats, and piglets. In vertebrates, C. sakazakii is a member of the normal (animal and human) oral and intestinal flora. It was found among the isolates from secretions of an infected mammary gland of dairy heifers. Liu et al. detected C. sakazakii in feed for pets. Also, C. sakazakii was isolated from various raw and ready-to-eat meat (products) [35, 43]. Isolated C. sakazakii has been found during a complicated curing process of meat products. It is a histamine-forming microorganism in the ripening process of cheese and has been isolated from a cheese whey substrate. Researchers demonstrated the lipolytic activity of a C. sakazakii-strain. C. sakazakii has been detected in fresh and prepared fish, too. A tetracycline-resistant C. sakazakii-strain from a Chilean freshwater salmon farm with no history of recent antibiotic use has been isolated. It has also been isolated from smoked sardines after 12 weeks of storage after irradiation [35, 44].
C. sakazakii with the total frequencies of 1.8% (10/564 strains) and 0.4% (1/256 strains) was detected when investigating the central and local drinking water supplies, respectively. During their investigation for biofilm formation, the bacteria indigenous to the water distribution system was found. Even bottled beverages should not be considered free of microorganisms [35, 45].
Transmission
Cronobacter sakazakii is thermo-tolerant and can contaminate PIF both intrinsically and extrinsically. Intrinsic contamination results from introducing the organism into the PIF at some stages during the manufacturing process. In contrast, extrinsic contamination may result from the use of contaminated utensils, such as blenders and spoons, to prepare PIF [46].
C. sakazakii are known to contaminate infant formula through the raw materials used for producing the formula or other dry ingredients after pasteurization; the caregiver may reconstitute the contamination of the formula just before feeding [47]. In most cases, the transmission route of Cronobacter-associated infant infections has not been confirmed. However, epidemiological studies in several incidents have implicated rehydrated PIF products as the vehicle of transmission and utensils used for formulae preparation as the likely source of contamination [19, 48].
Infection incidence
Although the number of well-documented cases of Cronobacter sakazakii infections in infants worldwide has increased in recent years, it has remained very low compared to many other infectious diseases. Also, there are reports in the literature about C. sakazakii (Cronobacter spp.) infections in older age groups, including adults, but the rate of infection among infants is lower. There has been no reported systematic review of cases in children and adults [49].
Pathogenicity and virulent factors
Virulence determinants of Cronobacter sakazakii are still unknown, and pathogenicity mechanisms have only begun to be researched. C. sakazakii is a foodborne pathogen that can cause severe illnesses and even death in persons with immunological deficiencies such as neonates, hospitalized patients in NICU, persons with severe underlying diseases, and the elderly. In these populations, this bacterium can successfully establish, colonize, and ultimately produce severe diseases [50, 51]. In humans, C. sakazakii has been found to affect specifically the nervous, gastrointestinal, and vascular systems. Establishment in the human vascular system causes bacteremia and or sepsis. It is often preceded by colonization beyond the blood-brain barrier leading to cerebrospinal fluid infection and meningitis, which may progress into intracerebral infarctions, brain abscess, and or cyst formation resulting in central nervous system deterioration. Necrotizing Enterocolitis (NEC) caused by C. sakazakii is currently the most common gastrointestinal emergency in neonates, characterized by necrosis of the gastrointestinal lumen [50-52].
Adhesions and attachment
Cronobacter sakazakii has adhesive capacities to several in vitro cell lines, including endothelial and transformed epithelial lines. This bacterium can attach to intestinal cells and survive in macrophages, but the specific involved receptors have remained to be determined. Recently, it was shown that the disruption of tight junctions significantly enhances the association of C. sakazakii with Caco-2 cells [53]. Some reports suggest a similarity between the tropism of Cronobacter sakazakii and Citrobacter koseri for invasion and infection of the central nervous system. It was noted that brain abscesses due to Cronobacter and Citrobacter koseri were morphologically similar and may be due to similar virulence mechanisms [53, 54].
Cronobacter sakazakii carries endotoxin on its surface; however, other virulence factors may also be crucial to pathogenicity. For a pathogen to cause an infection in a host, it should adhere to and colonize host surfaces. Specific adhesion to host cells is considered to be an essential virulence factor for most bacterial pathogens. This foodborne pathogen appears to adhere instantaneously to host surfaces and then proliferates in a logarithmic manner until an optimal concentration is attained. The adhesion of C.sakazakii to epithelial cells is mainly non-fimbriae-based, suggesting the role of other virulence factors in binding. The bacterium exhibits clustered adhesion, a pattern that also has been associated with neonatal colitis-causing strains of Klebsiella pneumoniae [28, 32]. C. sakazakii has an unusual surviving ability under dry conditions, but the thermal tolerance of C. sakazakii-strains may be different [32, 35].
Exposure to Cronobacter induced apoptosis and increased expression of interleukin-6 in infant rats could effectively explain the pathogenicity of Cronobacter-associated NEC in neonates [19].
The adhesive behavior of C. sakazakii strains is impressive, but until only recently, it has been thoroughly investigated. Adherence has been reported on inorganic and synthetic surfaces, such as plastics, silicon, PVC, polycarbonate, glass, stainless steel, and public water systems. Adhesion to living surfaces such as in a pathogenic capacity has been shown to differ between strains, but all share the trait of being independent of fimbriae structures and require a metabolically active host cell for adhesion [55].
Capsule
Some strains of Cronobacter sakazakii produce a viscous capsular material, potentially allowing the organism to form a biofilm on feeding equipment and contact surfaces, and how this material contributes to macrophage evasion remains to be determined [50, 56]. C. sakazakii capsule may also protect the organism, facilitating its survival in desiccated environments. The biofilm and capsule have been shown to reduce the efficacy of various common sanitizing methods, such as UV-light radiation, high osmotic pressures, heat, dry conditions, starvation, low pH, detergents, antibiotics, phagocytes, antibodies, and some bacteriophages. This biofilm may protect C. sakazakii, allowing it to survive osmotic, thermal, and tensile stressors [56]. Enterobacter sakazakii also produces a viscous capsular material, potentially allowing the organism to form a biofilm on feeding equipment and contact surfaces [38]. This biofilm may protect ES, allowing it to survive osmotic, thermal, and tensile stressors. Mange et al. demonstrated that ES has adhesive capacities to several in vitro cell lines, including endothelial and transformed epithelial lines [19].
Furthermore, entry is enhanced in the presence of the host cell cytoskeleton (actin filaments and microtubule structures) and disruption of the tight junction. This event may account for the opportunistic nature of this infection, particularly in infants and neonates [19].
The abilities of C. sakazakii to produce capsule and biofilm reduce the efficacy of UV-light, high osmotic pressure, heat, dry conditions, starvation, acids, inhibitory detergents like sanitizers and antibiotics, phagocytes, antibodies, and bacteriophages. C. sakazakii-strains can adhere to surfaces of materials and are used in the production, preparation, and administration of food like plastics, silicon, latex, polyvinyl chloride, less frequently on glass and stainless steel. C. sakazakii has been cultivated from biofilms of drinking water systems [35].
Outer membrane protein A (ompA)
Cronobacter sakazakii expresses the outer membrane protein A (ompA) that shows a high degree of homology with ompA genes of other Gram-negative bacteria [28]. The ompA protein is crucial in the invasion of brain endothelial cells by C. sakazakii. The organism also induces microtubule condensation at the sites of entry in endothelial cells. OmpA expression is required for moderate invasion of human intestinal epithelial cells. The binding of this bacterium to enterocytes, both in vitro and in the animal model, induces enterocyte apoptosis/necrosis in a dose-dependent manner [57].
Also, Cronobacter invades human brain microvascular endothelial cells more often than epithelial cells, but entry requires the expression of outer membrane protein A (ompA) to induce microtubule condensation [19].
Biofilm formation
Cronobacter sakazakii can attach to plastics and silicon rubber surfaces and grow in a biofilm. Enteral feeding tubes and feeding-bottle teats can harbor the bacterium in large numbers. Biofilm formation may also be a factor associated with altered susceptibility to antimicrobials [3].
Lipopolysaccharide
Large amounts of Lipopolysaccharide (LPS) are released by the bacteria killed by antibiotics, phagocytosis, the complement complex, or treatment with divalent cation chelators. In an infected host, small amounts of LPS can be protective by stimulating the immune system, while large amounts induce high fever and lead to septic shock and death by multiorgan failure and systemic inflammatory response. LPS liberated from bacteria is associated with LPS Binding Protein (LBP), an acute-phase protein present in the bloodstream, and forms complexes consisting of LPS, LBP, and soluble CD14 (sCD14) [58, 59]. The O-antigen region is one of the most variable regions in the membranes of Gram-negative bacteria, and this variability is routinely used to discriminate serotypes within the species of bacteria. Within O-antigen gene clusters, the wzx and wzy genes have the most diverse nucleotide sequences, which has led to their wide use in serotype-specific PCR assays [59, 60]. Finally, high levels of heat-stable lipopolysaccharide (endotoxin) in infant formula may enhance the translocation of Cronobacter across the gut and blood-brain barrier, increasing the risk of bacteremia in neonates [19, 58].
Viable but Nonculturable state (the VNC state)
Cronobacter sakazakii can become injured when they are subjected to stress such as heat and drying. This bacterium can respond to stress by entering a unique physiological state known as the “viable but nonculturable” (often abbreviated as the VNC state). Bacterial pathogens in the Enterobacteriaceae family (for example, Escherichia coli O157:H7, Salmonella, Shigella, Enterobacter cloacae, Klebsiella) can enter this state [17].
Minimum lethal dose
The actual amount of ES contamination usually is low, ranging from 0.36 to 66 Colony-Forming Units (CFU)/100 g [28].
Immune system state
In industrial countries, persons may have a predisposing condition (e.g. those receiving chemotherapy, transplant persons, people with chronic diseases, HIV positive patients, those receiving immunosuppressive treatments) that suppresses their T-cell mediated immunity, thereby potentially raising their susceptibility to a broad range of bacterial infections, including Cronobacter sakazakii infections. Also, an unknown proportion of neonates, infants, and toddlers may be temporarily immunocompromised at any time because of the pharmaceutical treatments, such as steroids for asthma exacerbations, injury, effects of acute disease, or stress. In addition, there has been a significant increase in the prescription of gastric acid-suppressing medications for gastroesophageal reflux in infants and toddlers in industrialized settings [61, 62]. While not traditionally considered immuno-suppressing, such medications impair one of the first lines of defense humans have against ingested pathogens. Infants, in addition, have lower levels of gastric acid, making them more vulnerable to infections. The prevalence of immunocompromising conditions may be relatively low in the developed countries, but the picture seems very different in developing countries, where the prevalence of such factors can rise to 40% [62].
Cronobacter sakazakii in powdered infant formula milk
Some of the vehicles of transmission and the infectious dose of Cronobacter sakazakii are unknown. The number of reported cases of Cronobacter infections is very low; nevertheless, it has slightly increased recently. Cronobacter sakazakii does not withstand the temperature at which milk is pasteurized, but it is easily found post-pasteurization, manufacturer environment, and preparation and handling before consumption. The thermal tolerance of Cronobacter in dried PIF has been well documented. This is why important steps towards preventive measures are needed to eliminate or avoid risks of PIF contamination and proliferation of the C. sakazakii. It is worth pointing out that breastfeeding should always be supported and encouraged since breast milk constitutes the preferred food for newborn infants, especially in their early months. When this is not possible, a mother should be well informed and educated on the importance of hygiene while handling, preparing, and storing PIF. Recommendations also underline the importance of using ready-to-feed PIF, complying with the rules for aseptic preparation and refrigeration at 2°C to 3°C of reconstituted PIF for a span of time shorter than 4 hours [2].
Treatment of infection caused by Cronobacter sakazakii
Traditionally, the treatment of Cronobacter infections with a combination of β-lactam (ampicillin) and aminoglycoside (gentamycin) or ampicillin-chloramphenicol has been successful [19]. It is sensitive to some antibiotics such as numerous β-lactams, antifolates, tetracycline, aminoglycosides, chloramphenicol, and quinolones. Also, trimethoprim-sulfamethoxazole may be useful. Nevertheless, many Cronobacter species are resistant to narrow-spectrum penicillins (i.e., ampicillin and amoxicillin) that traditionally have had good activity against members of the Enterobacteriaceae family. Emerging resistance of this bacterium to some antibiotics of choice should prompt physicians to consider carbapenems (ertapenem, meropenem, imipenem) or the newer broad-spectrum cephalosporins (e.g. cefepime) in combination with a second agent such as an aminoglycoside (e.g. gentamycin). Consequently, selecting antibiotics based on bacterial pure culture and susceptibility results and minimizing the use of broad-spectrum drugs are of paramount importance. C. sakazakii has intrinsic resistance to fusidic acids, streptogramins, glycopeptides, and lincosamides [28, 63].
Antimicrobial Resistance of Cronobacter sakazakii
C. sakazakii is intrinsically resistant to fosfomycin, rifampicin (rifampin), lincomycin, clindamycin, macrolides (clarithromycin, erythromycin, azithromycin), fusidic acid, and streptogramins. Nevertheless, resistance to ampicillin has increased owing to the production of β-lactamases and the acquisition of Transposable Elements (TEs) or jumping genes [2, 3, 64, 65]. Cronobacter sakazakii can inactivate cephalosporins and broad-spectrum antibiotics by secretion of β-lactamase enzymes [3, 64, 65].
Cronobacter sakazakii Resistance to routine sterilization methods
Cronobacter sakazakii is relatively resistant to dryness, heat, and osmotic stresses, which may explain, in part, its presence and survival in desiccated infant powder and similarly prepared products. Unsuitable storage and temperature regulation may raise the bacterial load, consequently facilitating infection outbreaks. Although C. sakazakii infection may increase secondary to reheating of formula and poor storage, it has not been documented definitely that staff of hospital themselves are not a vector for this foodborne pathogen. Unfortunately, hospital personnel may contribute to the spread of infection by ignoring suitable hygiene guidelines [66]. So, great care should be taken in contact with the isolation of infected and susceptible patients (infants) and adhering to strict hand washing. Also, C. sakazakii may form biofilms on different surfaces and, as a result of that acquiring resistance to disinfectants. It can survive in food powders for at least 12 months [28, 66, 67].
Tolerance or resistance of desiccated conditions for such a long time can be attributed to certain aspects of the Cronobacter sakazakii physiology, perhaps most notably because some isolates can produce a polysaccharide capsule. Over a 12-month storage period, the organism has been shown to survive better in dried formula products with an aw between 0.25 and 0.30 than at 0.69 and 0.82. C. sakazakii is resistant to desiccation over a wide range of aw (0.25-0.86) [68]. Sanitizer, disinfectants, and detergents reduce the C. sakazakii thermal resistance in powdered infant formula milk and other nutrient products.
Laboratory detection of Cronobacter sakazakii
An obstacle in diagnosing C. sakazakii infections and their sources is the ability or capacity of clinical, food, and environmental laboratories to identify the organism. The isolation of bacteria, such as C. sakazakii, from dried foods requires a series of steps to revitalize stressed cells that would otherwise not be cultured.
Cronobacter is a genus of the Enterobacteriaceae family. It is a Gram-negative, facultatively anaerobic, oxidase-negative, catalase-positive, and rod-shaped bacterium. They are generally motile, reduce nitrate, use citrate, hydrolyze esculin and arginine, and are positive for L-ornithine decarboxylation. Acid is produced from D-glucose, D-sucrose, D-raffinose, D-melibiose, D-cellobiose, D-mannitol, D-mannose, L-rhamnose, L-arabinose, D-trehalose, galacturonate and D-maltose. Cronobacter spp. are also generally positive for acetoin production (Voges–Proskauer test) and negative for the methyl red test, indicating 2,3-butanediol rather than mixed acid fermentation [68-71]. Some important biochemical tests to differentiate Cronobacter from Enterobacter spp. are presented in Table 1.

Isolation and identification from clinical samples
The detection of bacteria from typically sterile sites (e.g. blood, CSF) is less complex than their detection and isolation from a powdered formula, as the organisms would not be stressed and are unlikely to be in a mixed population. Nevertheless, accurate clinical identification of isolates such as C. sakazakii is restricted by using methods that have not been specifically validated for the organism [70].
Isolation and identification from food
Isolation of Cronobacter sakazakii from food samples is performed according to FDA protocol. Three Erlenmeyer flasks of sterile distilled water (pre-warmed to 45°C) of 9, 90, and 900 mL containing 1, 10, and 100 g of PIF, respectively, are prepared. After the weighed samples are thoroughly mixed and dissolved in distilled water, it was incubated at 35±2°C for 18 to 24 hours. After incubation, 10 mL of each sample was added to 90 mL of Enterobacteriaceae Enrichment (EE) broth medium and placed at 35±2°C for 18 to 24 hours. Following the incubation, a loopful of the enrichment culture is streaked onto duplicate Violet Red Bile Glucose Agar (VRBGA) medium plates, and incubation was performed for 18 to 24 hours at 35°C±2°C. Four presumptive colonies are picked from each VRBGA plate, and pure culture was performed on MacConkey agar and Tryptic Soy Agar (TSA). The isolated colonies that produce yellow pigment on TSA medium at room temperature (25°C) are identified. For the final confirmation of isolates, manual biochemical tests and biochemical tests embedded in the API-20E biochemical kit system are used [2].
Different media, including E. sakazakii Selective Broth (ESSB), Druggan-Forsythe-Iversen agar, Cronobacter Screening Broth (CSB), E. sakazakii Enrichment Broth (ESE), EE broth, and modified Lauryl Sulfate Broth (mLST), are used for the isolation of C. sakazakii from food samples. Compared with conventional, labor-intensive methods, the recently developed fluoro- and chromo-genic differential selective media decrease the time for the isolation of C. sakazakii [72]. The phenotypic identification of C. sakazakii with commercial systems may be inadequate for the strains isolated from food specimens. Molecular genetic methods revealed that several strains identified as C. sakazakii by commercial biochemical kits belonged to distinct species. To confirm C. sakazakii, more than one differentiation system is recommended. Computationally-based methods, including biochemical and 16S rDNA-data, improve the reliability and the detection time of identification [35, 73].
Molecular detection of Cronobacter sakazakii
Different techniques have been designed to identify Cronobacter sakazakii. Key molecular tests and particular areas of the DNA sequence can be applied for the accurate discrimination of C. sakazakii from other closely related species. To date, assays for the isolation and identification of C. sakazakii have applied production of yellow pigment and the α-glucosidase test as presumptive differentiating characteristics. However, these assays can result in presumptive false positives because of groups of as yet unidentified non-C. sakazakii Enterobacteriaceae, which are also positive for both of these characteristics. The use of yellow pigment as a defining characteristic can also result in false-negative because of the occurrence of non-pigmented C. sakazakii isolates and the occasional transient nature of this trait [5].
Prevention strategies
There are no active surveillance systems and guidelines for Cronobacter sakazakii disease. The World Health Organization and Food and Agriculture Organization of the United Nations made the following recommendations: (1) notify infant caregivers of the risks related to non-sterile powdered formula products, (2) encourage industry partners to develop a range of affordable sterile formula options, (3) if infants cannot breastfeed, consider feeding high-risk infants with sterile formula, and (4) consider setting an industry guideline for C. sakazakii and other Enterobacteriaceae members in infant formula milk. Factories warning labels on packages of powdered infant formula should stress that their products are non-sterile and need suitable handling, preparation, and storage and that sterile, liquid formula alternatives are available [26].
Pasteurization is effective in destroying Cronobacter sakazakii. Acidification reduced the concentration of C. sakazakii in vegetable-based foods and different types of infant formula products [35, 42]. Hygiene mismanagement due to incorrect time and temperature factors due to the transmission of bacteria via small vertebrates, insects, hands, and equipment should be avoided during the preparation process, production, and storage of food and drink. C. sakazakii may be related to food spoilage. It should be noted that the detection of the ubiquitous C. sakazakii in food is not always an indicator of hygiene mismanagement. For persons with acquired or congenital immunological deficiency, especially neonates, infants, elderly individuals, and persons with severe underlying diseases, the widespread occurrence of C. sakazakii and other Enterobacteriaceae members in the food and environment may indicate a health hazard [35].
2. Conclusion
Because of the ubiquitous nature of Cronobacter sakazakii and its unclear pathogenesis, preventive measures by parents, infant formula manufacturers, and health care providers will be necessary for the prevention of C. sakazakii-related infections. We recommend focusing on simple preventative strategies such as the promotion of breast milk feeding, the inclusion of warnings on powdered infant formula packages that they may be contaminated with C. sakazakii, and abstinence from the practice of re-warming of reconstituted formula. In adults, we believe that reconstituted dairy products should be avoided in immunosuppressed populations. Appropriate barrier precautions should be observed in ICU settings (both adult and neonatal), where the spread of infection may be more prevalent.
Ethical Considerations
Compliance with ethical guidelines
This article is a meta-analysis with no human or animal sample.
Funding
This study was supported financially by Gonabad University of Medical Sciences, Gonabad City, Iran.
Authors' contributions
All authors listed have contributed sufficiently to the project to be included as authors, and all those who are qualified to be authors are listed in the author byline.
Conflicts of interest
All authors declared no conflict of interest.
Acknowledgements
Our thanks go to Hassan Khajehei, PhD, for copy editing of the manuscript.
Refrences:
- Singh N, Goel G, Raghav M. Insights into virulence factors determining the pathogenicity of Cronobacter sakazakii. Virulence. 2015; 6(5):433-40. [DOI:10.1080/21505594.2015.1036217] [PMID] [PMCID]
- Mardaneh J, Soltan Dallal MM. Study of cronobacter sakazakii strains isolated from powdered milk infant formula by phenotypic and molecular methods in Iran. Archives of Pediatric Infectious Diseases. 2017; 5(1):e38867. [DOI:10.1128/JCM.42.11.5368-5370.2004] [PMID] [PMCID]
- Drudy D, Mullane NR, Quinn T, Wall PG, Fanning S. Enterobacter sakazakii: an emerging pathogen in powdered infant formula. Clinical Infectious Diseases. 2006; 42(7):996-1002. [DOI:10.1086/501019] [PMID]
- Iversen C, Waddington M, On SL, Forsythe S. Identification and phylogeny of enterobacter sakazakii relative to enterobacter and citrobacter species. Journal of Clinical Microbiology. 2004; 42(11):5368-70. [DOI:10.1128/JCM.42.11.5368-5370.2004] [PMID] [PMCID]
- Iversen C, Lancashire L, Waddington M, Forsythe S, Ball G. Identification of Enterobacter sakazakii from closely related species: The use of artificial neural networks in the analysis of biochemical and 16S rDNA data. BMC Microbiology. 2006; 6:28. [DOI:10.1186/1471-2180-6-28] [PMID] [PMCID]
- Leuschner RG, Bew J. A medium for the presumptive detection of Enterobacter sakazakii in infant formula: Inter laboratory study. Journal of AOAC International. 2004; 87(3):604-13. [DOI:10.1093/jaoac/87.3.604]
- Cai XQ, Yu HQ, Ruan ZX, Yang LL, Bai JS, Qiu DY, et al. Rapid detection and simultaneous genotyping of Cronobacter spp. (formerly Enterobacter sakazakii) in powdered infant formula using real-time PCR and high resolution melting (HRM) analysis. PLoS One. 2013; 8(6):e67082. [DOI:10.1371/journal.pone.0067082] [PMID] [PMCID]
- Iversen C, Lehner A, Mullane N, Marugg J, Fanning S, Stephan R, et al. Identification of ‘Cronobacter’ spp. (Enterobacter sakazakii). Journal of Clinical Microbiology. 2007; 45(11):3814-6. [DOI:10.1128/JCM.01026-07] [PMID] [PMCID]
- Dauga C, Breeuwer P. Taxonomy and physiology of Enterobacter sakazakii. In: Farber J, Forsythe S, Doyle M, editors. Enterobacter sakazakii. Washington D.C.: ASM Press; 2008. [DOI:10.1128/9781555815608.ch1] [PMID] [PMCID]
- Wikipedia. Cronus [Internet]. 2021 [Updated 2021 July 7]. Available from: https://en.wikipedia.org/wiki/Cronus
- FAO/WHO. Enterobacter sakazakii (Cronobacter spp.) in powdered follow-up formula: Microbiological risk assessment series 15. 1 Geneva, Switzerland; 2008. https://www.who.int/publications/i/item/9789241563796
- O’Brien S, Healy B, Negredo C, Anderson W, Fanning S, Iversen C. Prevalence of cronobacter species (enterobacter sakazakii) in follow-on infant formulae and infant drinks. Letters in Applied Microbiology. 2009; 48(5):536-41. [DOI:10.1111/j.1472-765X.2009.02562.x] [PMID]
- Park JH, Lee YD, Ryu TW. Chang HI. Identification and classification of cronobacter spp. isolated from powdered food in Korea. Journal of Microbiolog and Biotechnology. 2010; 20(4):757-62. [PMID]
- Zhao G, Yuan F, Yang H, Zhao Y, Chen Y. [Taxonomy of Enterobacter sakazakii and the biological characteristics of the new species and genus (Chinese)]. Wei sheng yan jiu= Journal of hygiene research. 2010; 39(2):248-50. [PMID]
- Parra-Flores J, Rodriguez A, Riffo F, Arvizu-Medrano SM, Arias-Rios EV, Aguirre J. Investigation on the factors affecting cronobacter sakazakii contamination levels in reconstituted powdered infant formula. Frontiers in Pediatrics. 2015; 3:72. [DOI:10.3389/fped.2015.00072] [PMID] [PMCID]
- Jongenburger I, Reij MW, Boer EP, Gorris LG, Zwietering MH. Actual distribution of cronobacter spp. in industrial batches of powdered infant formula and consequences for performance of sampling strategies. International Journal of Food Microbiology. 2011; 151(1):62-9. [DOI:10.1016/j.ijfoodmicro.2011.08.003] [PMID]
- Farmer JJ. My 40-year history with cronobacter/Enterobacter sakazakii-lessons learned, myths debunked, and recommendations. Frontiers in Pediatrics. 2015; 3:84. [DOI:10.3389/fped.2015.00084] [PMID] [PMCID]
- Iversen C, Lehner A, Mullane N, Bidlas E, Cleenwerck I, Marugg J, et al. The taxonomy of Enterobacter sakazakii: proposal of a new genus Cronobacter gen. nov. and descriptions of Cronobacter sakazakii comb. nov. Cronobacter sakazakii subsp. sakazakii, comb. nov., Cronobacter sakazakii subsp. malonaticus subsp. nov., Cronobacter turicensis sp. nov., Cronobacter muytjensii sp. nov., Cronobacter dublinensis sp. nov. and Cronobacter genomospecies 1. BMC Evolutionary Biology. 2007; 7:64. [DOI:10.1186/1471-2148-7-64] [PMID] [PMCID]
- Chenu JW, Cox JM. Cronobacter (‘Enterobacter sakazakii’): current status and future prospects. Letters in Applied Microbiology. 2009; 49(2):153-9. [DOI:10.1111/j.1472-765X.2009.02651.x] [PMID]
- Iversen C, Mullane N, McCardell B, Tall BD, Lehner A, Fanning S, et al. Cronobacter gen. nov., a new genus to accommodate the biogroups of Enterobacter sakazakii, and proposal of Cronobacter sakazakii gen. nov., comb. nov., Cronobacter malonaticus sp. nov., Cronobacter turicensis sp. nov., Cronobacter muytjensii sp. nov., Cronobacter dublinensis sp. nov., Cronobacter genomospecies 1, and of three subspecies, Cronobacter dublinensis subsp. dublinensis subsp. nov., Cronobacter dublinensis subsp. lausannensis subsp. nov. and Cronobacter dublinensis subsp. lactardi subsp. International Journal of Systematic and Evolutionary Microbiology. 2008; 58(Pt 6):1442-7. [DOI:10.1099/ijs.0.65577-0] [PMID]
- Kim KP, Klumpp J, Loessner MJ. Enterobacter sakazakii bacteriophages can prevent bacterial growth in reconstituted infant formula. International Journal of Food Microbiology. 2007; 115(2):195-203. [DOI:10.1016/j.ijfoodmicro.2006.10.029] [PMID]
- Davin-Regli A, Lavigne JP, Pagès JM. Enterobacter spp.: update on taxonomy, clinical aspects, and emerging antimicrobial resistance. Clinical Microbiology Reviews. 2019; 32(4):e00002-19. [DOI:10.1128/CMR.00002-19]
- Iversen C, Lane M, Forsythe SJ. The growth profile, thermotolerance and biofilm formation of Enterobacter sakazakii grown in infant formula. Letters in Applied Microbiology. 2004; 38(5):378-82. [DOI:10.1111/j.1472-765X.2004.01507.x] [PMID]
- Mittal R, Wang Y, Hunter CJ, Gonzalez-Gomez I, Prasadarao NV. Brain damage in newborn rat model of meningitis by Enterobacter sakazakii: a role for outer membrane protein A. Laboratory Investigation. 2009; 89(3):263-77. [DOI:10.1038/labinvest.2008.164] [PMID] [PMCID]
- Block C, Peleg O, Minster N, Bar-Oz B, Simhon A, Arad I, et al. Cluster of neonatal infections in Jerusalem due to unusual biochemical variant of Enterobacter sakazakii. European Journal of Clinical Microbiology & Infectious Diseases. 2002; 21(8):613-6. [DOI:10.1007/s10096-002-0774-5] [PMID]
- Bowen AB, Braden CR. Invasive Enterobacter sakazakii disease in infants. Emerging Infectious Diseases. 2006; 12(8):1185-9. [DOI:10.3201/eid1208.051509] [PMID] [PMCID]
- Centers for Disease Control Prevention. Enterobacter sakazakii infections associated with the use of powdered infant formula-Tennessee, 2001. MMWR. Morbidity and Mortality Weekly Report. 2002; 51(14):297-300. [PMID]
- Hunter CJ, Petrosyan M, Ford HR, Prasadarao NV. Enterobacter sakazakii: An emerging pathogen in infants and neonates. Surgical Infections. 2008; 9(5):533-9. [DOI:10.1089/sur.2008.006] [PMID] [PMCID]
- Healy B, Cooney S, O'Brien S, Iversen C, Whyte P, Nally J, et al. Cronobacter (Enterobacter sakazakii): An opportunistic foodborne pathogen. Foodborne Pathogens and Disease. 2010; 7(4):339-50. [DOI:10.1089/fpd.2009.0379] [PMID]
- Cho TJ, Hwang JY, Kim HW, Kim YK, Il Kwon J, Kim YJ, et al. Underestimated risks of infantile infectious disease from the caregiver's typical handling practices of infant formula. Scientific Reports. 2019; 9(1):9799. [DOI:10.1038/s41598-019-46181-0] [PMID]
- Jackson EE, Flores JP, Fernández-Escartín E, Forsythe SJ. Re-evaluation of a suspected Cronobacter sakazakii outbreak in Mexico. Journal of Food Protection. 2015; 78(6):1191-6. [DOI:10.4315/0362-028X.JFP-14-563] [PMID]
- Gurtler JB, Beuchat LR. Inhibition of growth of Enterobacter sakazakii in reconstituted infant formula by the lactoperoxidase system. Journal of Food Protection. 2007; 70(9):2104-10. [DOI:10.4315/0362-028X-70.9.2104] [PMID]
- Hurrell E, Kucerova E, Loughlin M, Caubilla-Barron J, Hilton A, Armstrong R, Smith C, Grant J, Shoo S, Forsythe S. Neonatal enteral feeding tubes as loci for colonisation by members of the Enterobacteriaceae. BMC Infect Dis. 2009; 9(1):146. [DOI:10.1186/1471-2334-9-146]
- Lai KK. Enterobacter sakazakii infections among neonates, infants, children, and adults. Case reports and a review of the literature. Medicine. 2001; 80(2):113-22. [DOI:10.1097/00005792-200103000-00004] [PMID]
- Friedemann M. Enterobacter sakazakii in food and beverages (other than infant formula and milk powder). International Journal of Food Microbiology. 2007; 116(1):1-10. [DOI:10.1016/j.ijfoodmicro.2006.12.018] [PMID]
- Kandhai MC, Reij MW, Gorris LG, Guillaume-Gentil O, van Schothorst M. Occurrence of Enterobacter sakazakii in food production environments and households. Lancet. 2004; 363(9402):39-40. [DOI:10.1016/S0140-6736(03)15169-0]
- O’Hara CM, Miller JM. Evaluation of the Vitek 2 ID-GNB assay for identification of members of the family Enterobacteriaceae and other nonenteric gram-negative bacilli and comparison with the Vitek GNI+ card. Journal of Clinical Microbiology. 2003; 41(5):2096-101. [DOI:10.1128/JCM.41.5.2096-2101.2003] [PMID] [PMCID]
- Breeuwer P, Lardeau A, Peterz M, Joosten HM. Desiccation and heat tolerance of Enterobacter sakazakii. Journal of Applied Microbiology. 2003; 95(5):967-73. [DOI:10.1046/j.1365-2672.2003.02067.x] [PMID]
- Lehner A, Tasara T, Stephan R. 16S rRNA gene based analysis of Enterobacter sakazakii strains from different sources and development of a PCR assay for identification. BMC Microbiology. 2004; 25; 4:43. [DOI:10.1186/1471-2180-4-43] [PMID] [PMCID]
- Kanivets VI, Pishchur IN. Bacterial microflora on disinfected sugarbeet seeds. Microbiology. 2001; 70(3):316-8. [DOI:10.1023/A:1010407528520]
- Kuklinsky-Sobral J, Araujo WL, Mendes R, Pizzirani-Kleiner AA, Azevedo JL. Isolation and characterization of endophytic bacteria from soybean (Glycine max) grown in soil treated with glyphosate herbicide. Plant and Soil. 2005; 273:91-9. [DOI:10.1007/s11104-004-6894-1]
- Coulin P, Farah Z, Assanvo J, Spillmann H, Puhan Z. Characterization of the microflora of attieke, a fermented cassava product, during traditional small-scale preparation. International Journal of Food Microbiology. 2006; 106(2):131-6. [DOI:10.1016/j.ijfoodmicro.2005.06.012] [PMID]
- Liu Y, Cai X, Zhang X, Gao Q, Yang X, Zheng Z, et al. Real time PCR using TaqMan and SYBR green for detection of Enterobacter sakazakii in infant formula. Journal of Microbiological Methods. 2006; 65(1):21-31. [DOI:10.1016/j.mimet.2005.06.007] [PMID]
- Chaves-Lopez C, De Angelis M, Martuscelli M, et al. Characterization of the Enterobacteriaceae isolated from anartisanal Italian ewe’s cheese (Pecorino Abruzzese). Journal of Applied Microbiology. 2006; 101:353-60. [DOI:10.1111/j.1365-2672.2006.02941.x] [PMID]
- Lee DG, Kim SJ. Bacterial species in biofilm cultivated from the end of the Seoul water distribution system. Journal of Applied Microbiology. 2003; 95(2):317-24. [DOI:10.1046/j.1365-2672.2003.01978.x] [PMID]
- Ling N, Forsythe S, Wu Q, Ding Y, Zhang J, Zeng H. Insights into Cronobacter sakazakii biofilm formation and control strategies in the food industry. Engineering. 2020; 6(4):393-405. [DOI:10.1016/j.eng.2020.02.007]
- Casalinuovo F, Rippa P, Battaglia L, Parisi N. Isolation of Cronobacter spp. (Enterobacter sakazakii) from Artisanal Mozzarella. Italian Journal of Food Safety. 2014; 3(1):1526. [DOI:10.4081/ijfs.2014.1526] [PMID] [PMCID]
- Caubilla-Barron J, Hurrell E, Townsend S, Cheetham P, Loc-Carrillo C, Fayet O, et al. Genotypic and phenotypic analysis of Enterobacter sakazakii strains from an outbreak resulting in fatalities in a neonatal intensive care unit in France. Journal of Clinical Microbiology. 2007; 45(12):3979-85. [DOI:10.1128/JCM.01075-07] [PMID] [PMCID]
- Camacho-Gonzalez A, Spearman PW, Stoll BJ. Neonatal infectious diseases: evaluation of neonatal sepsis. Pediatric Clinics of North America. 2013; 60(2):367-89. [DOI:10.1016/j.pcl.2012.12.003] [PMID] [PMCID]
- Singh VP. Recent approaches in food bio-preservation - a review. Open Vet J. 2018; 8(1):104-111. [DOI:10.4314/ovj.v8i1.16]
- Mange JP, Stephan R, Borel N, Wild P, Kim KS, Pospischil A, et al. Adhesive properties of Enterobacter sakazakii to human epithelial and brain microvascular endothelial cells. BMC Microbiology. 2006; 6:58. [DOI:10.1186/1471-2180-6-58] [PMID] [PMCID]
- Mullane NR, Whyte P, Wall PG, Quinn T, Fanning S. Application of pulsed-field gel electrophoresis to characterize and trace the prevalence of Enterobacter sakazakii in an infant formula processing facility. International Journal of Food Microbiology. 2007; 116(1):73-81. [DOI:10.1016/j.ijfoodmicro.2006.12.036] [PMID]
- Kim KP, Loessner MJ. Enterobacter sakazakii invasion in human intestinal Caco-2 cells requires the host cell cytoskeleton and is enhanced by disruption of tight junction. Infection and Immunity. 2008; 76(2):562-70. [DOI:10.1128/IAI.00937-07] [PMID] [PMCID]
- Kothary MH, Gopinath GR, Gangiredla J, Rallabhandi PV, Harrison LM, Yan QQ, et al. Analysis and characterization of proteins associated with outer membrane vesicles secreted by cronobacter spp. Frontiers in Microbiology. 2017; 8:134. [DOI:10.3389/fmicb.2017.00134] [PMID] [PMCID]
- Leclercq A, Wanegue C, Baylac P. Comparison of fecal coliform agar and violet red bile lactose agar for fecal coliform enumeration in foods. Appl Environ Microbiol. 2002; 68(4):1631-8. [DOI:10.1128/AEM.68.4.1631-1638.2002]
- Kim H, Ryu JH, Beuchat LR. Effectiveness of disinfectants in killing Enterobacter sakazakii in suspension, dried on the surface of stainless steel, and in a biofilm. Applied and Environmental Microbiology. 2007; 73(4):1256-65. [DOI:10.1128/AEM.01766-06] [PMID] [PMCID]
- Mohan Nair MK, Venkitanarayanan KS. Cloning and sequencing of the ompA gene of Enterobacter sakazakii and development of an ompA-targeted PCR for rapid detection of Enterobacter sakazakii in infant formula. Applied and Environmental Microbiology. 2006; 72(4):2539-46. [DOI:10.1128/AEM.72.4.2539-2546.2006] [PMID] [PMCID]
- Rosenfeld Y, Shai Y. Lipopolysaccharide (Endotoxin)-host defense antibacterial peptides interactions: Role in bacterial resistance and prevention of sepsis. Biochimica et Biophysica Acta. 2006; 1758(9):1513-22. [DOI:10.1016/j.bbamem.2006.05.017] [PMID]
- Matsuura M. Structural modifications of Bacterial lipopolysaccharide that facilitate gram-negative Bacteria evasion of host innate Immunity. Frontiers in Immunology. 2013; 4:109. [DOI:10.3389/fimmu.2013.00109] [PMID] [PMCID]
- Jarvis KG, Grim CJ, Franco AA, Gopinath G, Sathyamoorthy V, Hu L, et al. Molecular characterization of Cronobacter lipopolysaccharide O-antigen gene clusters and development of serotype-specific PCR assays. Applied and Environmental Microbiology. 2011; 77(12):4017-26. [DOI:10.1128/AEM.00162-11] [PMID] [PMCID]
- Khoshoo V, Edell D, Thompson A, Rubin M. Are we overprescribing antireflux medications for infants with regurgitation? Pediatrics. 2007; 120(5):946-9. [DOI:10.1542/peds.2007-1146] [PMID]
- Savino F, Castagno E. Over prescription of antireflux medications for infants with regurgitation. Pediatrics. 2008; 121(5):1070. [DOI:10.1542/peds.2008-0179] [PMID]
- Ray P, Das A, Gautam V, Jain N, Narang A, Sharma M. Enterobacter sakazakii in infants: novel phenomenon in India. Indian JMed Microbiol. 2007; 25(4):408. [DOI:10.4103/0255-0857.37351]
- Stock I, Wiedemann B. Natural antibiotic susceptibility of Enterobacter amnigenus, Enterobacter cancerogenus, Enterobacter gergoviae and Enterobacter sakazakii strains. Clinical Microbiology and Infection. 2002; 8(9):564-78. [DOI:10.1046/j.1469-0691.2002.00413.x] [PMID]
- Girlich D, Poirel L, Leelaporn A, Karim A, Tribuddharat C, Fennewald M, et al. Molecular epidemiology of the integron-located VEB-1 extended-spectrum beta-lactamase in nosocomial enterobacterial isolates in Bangkok, Thailand. Journal of Clinical Microbiology. 2001; 39(1):175-82. [DOI:10.1128/JCM.39.1.175-182.2001] [PMID] [PMCID]
- Gould DJ, Moralejo D, Drey N, Chudleigh JH, Taljaard M. Interventions to improve hand hygiene compliance in patient care. Cochrane Database of Systematic Reviews. 2017; 9(9):CD005186. [DOI:10.1002/14651858.CD005186.pub4] [PMID] [PMCID]
- Bar-Oz B, Preminger A, Peleg O, Block C, Arad I. Enterobacter sakazakii infection in the newborn. Acta Paediatr. 2001; 90(3):356-8. [PMID]
- Kent RM, Fitzgerald GF, Hill C, Stanton C, Ross RP. Novel approaches to improve the intrinsic microbiological safety of powdered infant milk formula. Nutrients. 2015; 7(2):1217-44. [DOI:10.3390/nu7021217] [PMID] [PMCID]
- Farmer JJ III. Enterobacteriaceae: introduction and identification. In: MurrayPR, editors. Manual of cinical microbiology. 7th ed. Washington DC: ASM Press Inc; 1999. https://www.worldcat.org/title/manual-of-clinical-microbiology/oclc/39914150
- Stephan R, Van Trappen S, Cleenwerck I, Vancanneyt M, De Vos P, Lehner A. Enterobacter turicensis sp. nov. and Enterobacter helveticus sp. nov., isolated from fruit powder. International Journal of Systematic and Evolutionary Microbiology. 2007; 57(Pt 4):820-6. [DOI:10.1099/ijs.0.64650-0] [PMID]
- Gebremariam A. Neonatal meningitis in Addis Ababa: A 10-year review. Ann Trop Paediatr. 1998; 18(4):279-83. [DOI:10.1080/02724936.1998.11747960]
- Restaino L, Frampton EW, Lionberg WC, Becker RJ. A chromogenic plating medium for the isolation and identification of Enterobacter sakazakii from foods, food ingredients, and environmental sources. Journal of Food Protection. 2006; 69(2):315-22. [DOI:10.4315/0362-028X-69.2.315] [PMID]
- Ball G, Mian S, Holding F, Allibone RO, Lowe J, Ali S, et al. An integrated approach utilizing artificial neural networks and SELDI mass spectrometry for the classification of human tumours and rapid identification of potential biomarkers. Bioinformatics. 2002; 18(3):395-404.[DOI:10.1093/bioinformatics/18.3.395]
Type of Study: Review |
Subject:
Laboratory Medicine
Received: 2019/11/25 | Accepted: 2022/03/13 | Published: 2021/04/1
Received: 2019/11/25 | Accepted: 2022/03/13 | Published: 2021/04/1
| Rights and permissions | |
 |
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License. |





